Edge AI Anomaly Detection Part 2 - Feature Extraction and Model Training
2020-05-18 | By ShawnHymel
License: Attribution
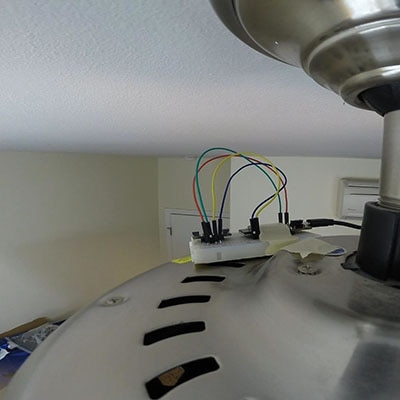
In the previous tutorial, we collected vibration data from a ceiling fan using an ESP32 and accelerometer. This time, we perform various mathematical calculations and transforms on the data to look for features that can help us discern an anomalous condition from a normal condition. Once we have chosen our features, we will train two different machine learning models: a more classical model involving the Mahalanobis Distance and another using a neural network.
You can watch this tutorial in video format here:
Feature Extraction
In most machine learning systems, we cannot or do not want to send raw data directly to our model. Some models, like neural networks, can be trained to extract features, but it often comes with increased computational complexity. If we can determine which features will allow us to detect anomalies (or predict values or classify cases) most easily, it will save lots of training effort and processing power down the road.
A “feature” can be almost anything that is extracted from the raw data. It could be the raw data itself, combinations of different sensor data, statistical analysis on several measurements (mean, variance, etc.), or transformations, such as the Fourier transform to the frequency domain. Once we’ve picked out one or more features, we can then train a model using those features. From then on, whenever we want to use the model to make predictions/classifications, we must extract the same features from new data.
There are many algorithms that can be used to automate the process of feature extraction and reduce the number of dimensions going into a machine learning model. For images, this might be something like edge detection. For sounds, this might be a fast Fourier transform (FFT). In other cases, you might see something like the discrete cosine transform (DCT). Algorithms, like k-means clustering and principal component analysis (PCA) can help determine groupings and reduce dimensions in your data.
We will not get into these algorithms in this tutorial. Instead, we are going to look at several simpler statistical analysis tools, like variance, and use visual inspection to determine if something makes for a good feature.
Dataset
You are welcome to collect your own data by following the steps outlined in the previous tutorial. If you would like to use the data that I collected from my ceiling fan, download this repository and look in the ceiling-fan-dataset directory for .csv files containing raw accelerometer data (units of g-force).
Analyze Raw Data
In a new Jupyter Notebook, run the code found here: https://github.com/ShawnHymel/tinyml-example-anomaly-detection/blob/master/data_collection/anomaly-detection-feature-analysis.ipynb
It will read data from the files stored in the fan_0_low_0_weight folder and compare it to all other settings. For now, we will assume that fan low, no weights added (coins taped to the blades) is our “normal” condition. Anything that deviates from that will be considered an “anomaly.”
Since we are interested in vibration data, we need to subtract out the mean of each sample set from every measurement in that set. This effectively removed the “DC” component so that we only look at variations. If the accelerometer was positioned at one angle or slightly moved during data collection, you would see that show up in one of the axes. Removing the DC component helps combat that.
Data scientists often like to look at several different statistical properties when dealing with groups of data: mean, variance, kurtosis, and skew. We’ll plot each of these and a couple others.
Note that each dot on the following plots represents some statistical analysis (mean, variance, etc.) performed on one file with 200 measurements. So, the mean_x is the average of all the measurements in the x axis, the mean_y is the average of all the measurements in the y axis, and the mean_z is the average of all the measurements in the z axis.
Blue dots are from fan low, no weights (normal) and red dots are from everything else (anomaly).
First, we look at all of the means, which should be mostly useless at this point. Subtracting the mean from every sample means most samples should appear around 0.
Next, we want to look at variance in each data set, where variance is the squared deviation from the mean.
Do you see how the group of blue dots is somewhat separate from the red dots? This separation is important: with separation between groups, we can easily draw a boundary between them. This is where we get into machine learning to discern the difference between groups like this.
Let’s keep going. Kurtosis tells us how much of a tail a distribution has (relative to a Gaussian/normal distribution).
There’s some separation between blue and red dots here, but most of them are mixed up around (0, 0, 0). It would be difficult to draw a clear boundary between them.
Skew (or skewness) tells us how symmetrical (or asymmetrical) a distribution is.
It looks potentially useful. If you are using an interactive matplotlib session on Jupyter Notebook, you can move the plots around in 3D. The skew, unfortunately, does seem to be a great predictor of anomalies, either. You can read more about kurtosis and skew here.
Next, we have median absolute deviation (MAD). While standard deviation (and variance) are great at describing the spread of data in a normal distribution, they can easily be affected by outliers and non-normally distributed data. As a result, MAD offers a more robust way to measure spread for non-normal data. Since you’ll often find data that is not normally distributed, MAD can be the best feature to measure spread.
As we can see, the blue dots are clearly separated from the red dots, which indicates that MAD is a fantastic way to discern normal operation from other vibration data.
Sometimes, correlation offers insights into how data is related along its various axes. If we plot the histograms comparing the correlation between each set of axes, we can see if there’s any separation between the normal and anomalous samples.
There seems to be some separation on the plots, but not nearly as much as MAD.
Finally, we can look at the frequency domain (via the FFT) to see if that offers any insights. Note that bin 0 (DC and low frequencies) has been removed so we can view the higher frequencies more easily in the following plot.
There might be some information we can glean from this that would lead to possible anomaly detections, but in a resource limited device (such as a microcontroller), computing the FFT can be expensive. And once again, the MAD offers a much cleaner separation between normal and anomaly features.
Machine Learning Model 1: Mahalanobis Distance
For each sample (consisting of 200 measurements each), we calculate the MAD in the X axis, MAD in the Y axis, and MAD in the Z axis. We can plot the MAD values (x, y, z) for each sample on a 3D grid, as we showed in the previous section.
Let’s first look at finding an outlier using 2 dimensions instead of 3 (as it makes things a little easier to visualize). If it helps, imagine looking at just the X and Y axes and leave out Z for now. Note that the same principles apply for finding outliers with any number of dimensions.
For “normal” operation, these MAD values should form a cluster, ideally away from other clusters. Our normal operation cluster might look something like this (simplifying to 2D by removing out the Z axis):
This shows that X and Y are uncorrelated. As such, we can find the mean of the cluster (denoted by the black ‘+’ symbol) and set a boundary around the cluster. We then introduce a new sample. If the distance between the mean of the cluster and the new sample is greater than some threshold, we say the new sample is an “anomaly.” If not, then it’s classified as “normal.”
The distance measured here is the simple Euclidean distance.
Unfortunately, you’ll often find that data is correlated, like in the case of our accelerometer readings. If we were to use the Euclidean distance between a new point and mean (with correlated data), we’d run into a problem:
The red dot looks like an anomaly to our eyes, but if we use the Euclidean distance, it would be considered within the threshold and therefore, a normal point.
To fix that, we turn to the Mahalanobis distance. It takes principal components into account, which allows us to draw an ellipse as a boundary instead of a circle (I won’t get into the math here, but you can read more about the Mahalanobis distance here):
When we calculate the Mahalanobis distance, we get a single number, so it’s easy to compare to a threshold. Anything over that threshold is an anomaly, and anything equal to or less than it is considered normal.
In our Python training program, we just need to read from the CSV files and find the mean and covariance matrices of the “normal” samples (note that normal samples are from the fan_0_low_0_weight folder). See the Python code here to dig into it further.
Once we have the mean and covariance matrix, we can then write a function that calculates the Mahalanobis distance between the mean and a new point:
# Calculate mahalanobis distance of x from group described by mu, cov # Based on: https://www.machinelearningplus.com/statistics/mahalanobis-distance/ def mahalanobis(x, mu, cov): x_minus_mu = x - mu inv_covmat = sp.linalg.inv(cov) left_term = np.dot(x_minus_mu, inv_covmat) mahal = np.dot(left_term, x_minus_mu.T) if mahal.shape == (): return mahal else: return mahal.diagonal()
We test it out by calculating the Mahalanobis distance with all normal and anomaly samples and plot the histograms of their distances (blue is normal distances, red is anomaly distances).
As you can see, there is a clear separation between the red and blue groupings. All of the normal samples are less than about 20 for their Mahalanobis distance whereas the anomaly samples are above that.
Using this, we can create a simple threshold. Any new sample whose Mahalanobis distance is above 20 is an “anomaly” and anything else is “normal.” We can use our test set that we put aside at the beginning of the program to generate this confusion matrix:
With that, we can see that setting a threshold of 20 creates a perfect division for our test set! We would be ready to deploy our model (mean and covariance matrices). Note that this model only describes one particular fan at one particular speed. We would need a lot more data to characterize all ceiling fans or we would want to create a unique model for each fan.
We can save the model by storing the mean and covariance matrices in a .npz file.
Machine Learning Model 2: Autoencoder Neural Network
The Mahalanobis distance is likely faster and more robust, but since neural networks are all the rage right now, let’s make an anomaly detection system using them! A popular way to configure a neural network to detect anomalies is the “autoencoder.”
An autoencoder consists of several layers of nodes. The first few layers shrink in node size (the encoder) and the second set of layers increase in node size (the decoder). The basic idea behind an autoencoder is that it attempts to recreate whatever was given to it at the input layer.
For example, an autoencoder trained to work with images would find features in the encoder section and then attempt to recreate the image in the decoder section. So, your output should look something like the input.
We can measure how well the autoencoder performs by calculating the mean squared error (MSE) between the input values and predicted (output) values (i.e. how well did the autoencoder do at recreating the input values/image).
To use an autoencoder for anomaly detection, we train the autoencoder on only the normal samples. If done right, the MSE for any new normal samples should be low, as the autoencoder should be able to figure out the relationships and features necessary for reproducing the same MAD values as the input MAD values.
However, if given anomaly MAD values, the autoencoder should struggle to reproduce the same values, resulting in a higher MSE between the input and output. With the right neural network model and training, we should see low MSE values for normal samples and high MSE values for anomalies.
After playing around with some different neural network architectures, I found that a simple 2-layer network worked well and kept the number of calculations small (necessary for running a neural network on a microcontroller!)
The code for building and training this autoencoder can be found here.
If we plot the normal versus anomaly MSEs (as histograms), we should see some separation, denoting that the autoencoder can recreate the normal MAD samples easier than the anomaly MAD samples. I recommend trying this with the validation data.
Note that if you do not see a clear separation between the normal and anomaly MSEs (blue and red), you should try running the Jupyter Notebook cells that create the model and train the model again until you do (training weights are randomly initialized, which can affect how well the autoencoder works). For something in production, you might want to automate this task by putting the model training in a for-loop that looks for the smallest MSE value on the training data (normal samples).
Just like in the Mahalanobis distance example, we can create a simple classifier to use on our test set and create a confusion matrix to determine how well our autoencoder performs at detecting anomalies.
Once we’re happy with the performance of the model, we can save it in a Keras .h5 file.
Resources and Going Further
One thing we did not cover in this demonstration is dimension reduction. Reducing the number of dimensions is crucial in reducing the size and complexity of the model (including neural networks). One of the main techniques for dimension reduction is principal component analysis (PCA). This article offers a great visual introduction to PCA: https://setosa.io/ev/principal-component-analysis/
If you wish to learn more about the Mahalanobis distance, this StackExchange thread has a great explanation: https://stats.stackexchange.com/questions/62092/bottom-to-top-explanation-of-the-mahalanobis-distance
Finally, see this article if you wish to read more about autoencoders (and an example used to detect credit card fraud): https://medium.com/@curiousily/credit-card-fraud-detection-using-autoencoders-in-keras-tensorflow-for-hackers-part-vii-20e0c85301bd
Recommended Reading
Edge AI Anomaly Detection Part 3 - Machine Learning on Raspberry Pi
Edge AI Anomaly Detection Part 4 - Machine Learning on ESP32 via Arduino